Vennt
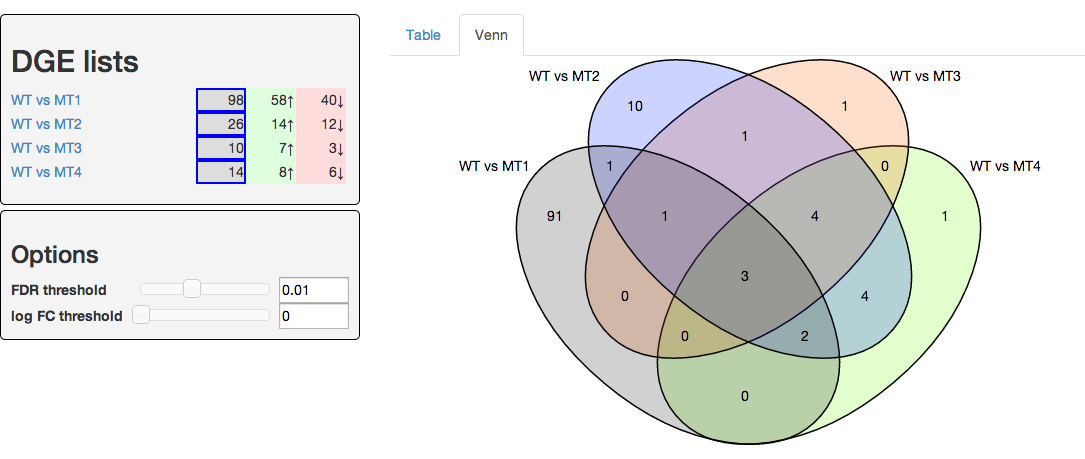
A web-tool to generate dynamic Venn diagrams for differential gene expression
Try a live demoCreating your own
Download this python script vennt.py (requires python >=2.7).
Using Cuffdiff output
If you have used Cuffdiff to perform your differential expression analysis, then you can use a simple one-liner to generate an html file for your results. Run:
python vennt.py --cuffdiff gene_exp.diff > my-vennt.html
This will create a single html file that can be shared.
Using a general CSV file of DGE
The output from other differential expression software can also be used with Vennt. We commonly use Voom/Limma and edgeR. Generate a CSV file containing your gene-lists, formatted like the following:
key,Feature,Description,Gene Name,logFC,adj.P.Val
WT vs MT1,ENSG00000083520,DIS3 mitotic control homolog (S. cerevisiae),DIS3,-2.4,4.8e-10
WT vs MT1,ENSG00000025156,heat shock transcription factor 2,HSF2,-0.89,6.4e-05
WT vs MT1,ENSG00000103042,"solute carrier family 38, member 7",SLC38A7,1.5,6.4e-05
WT vs MT2,ENSG00000083520,DIS3 mitotic control homolog (S. cerevisiae),DIS3,-2.4,4.8e-10
WT vs MT2,ENSG00000025156,heat shock transcription factor 2,HSF2,-0.89,6.4e-05
WT vs MT2,ENSG00000103042,"solute carrier family 38, member 7",SLC38A7,1.5,6.4e-05
Then, if your CSV column names match the defaults, simply run it as follows:
python vennt.py gene-lists.csv > my-vennt.html
You may need to configure the column names used in your CSV file. This is done using arguments to vennt.py (read the command line help).
It is also possible to use separate CSV files for your gene lists. Each CSV file is expected to be in the same format with columns in the same order. The 'key' column is generated for you using the names of the files you specify. To use separate CSV files simply specfiy them on the command line to vennt.py.
Separate HTML and CSV file
As an alternative to embedding the CSV file, you can have it served from a web server. Simply download this html (right-click 'Save As'), then put it and your CSV file on a webserver.
You can configure some settings in that html file, such as the CSV filename, or your column names. Read the documentation for a complete description.